How scientists are turning millions of underwater images
into usable reef knowledge, faster.
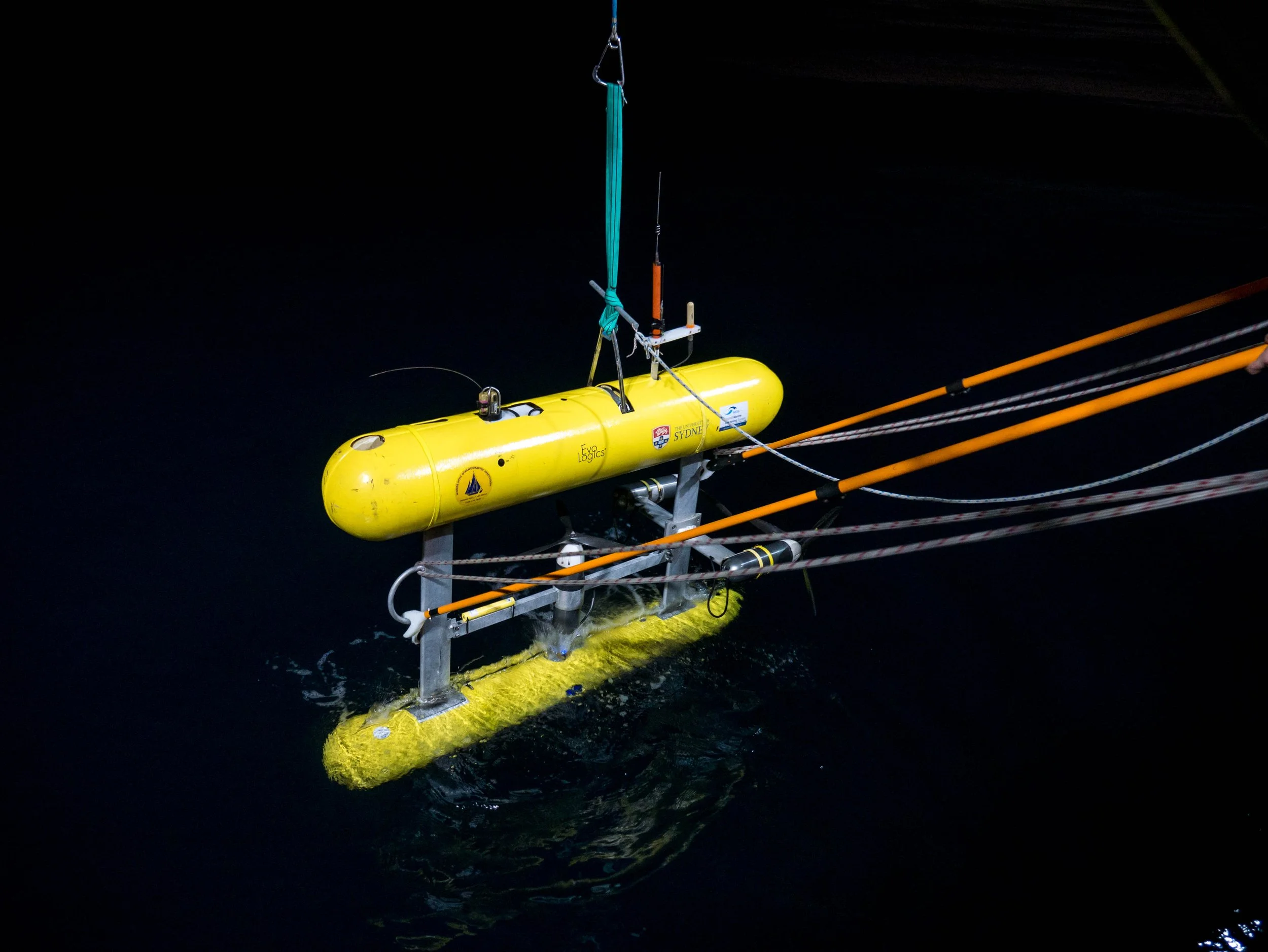

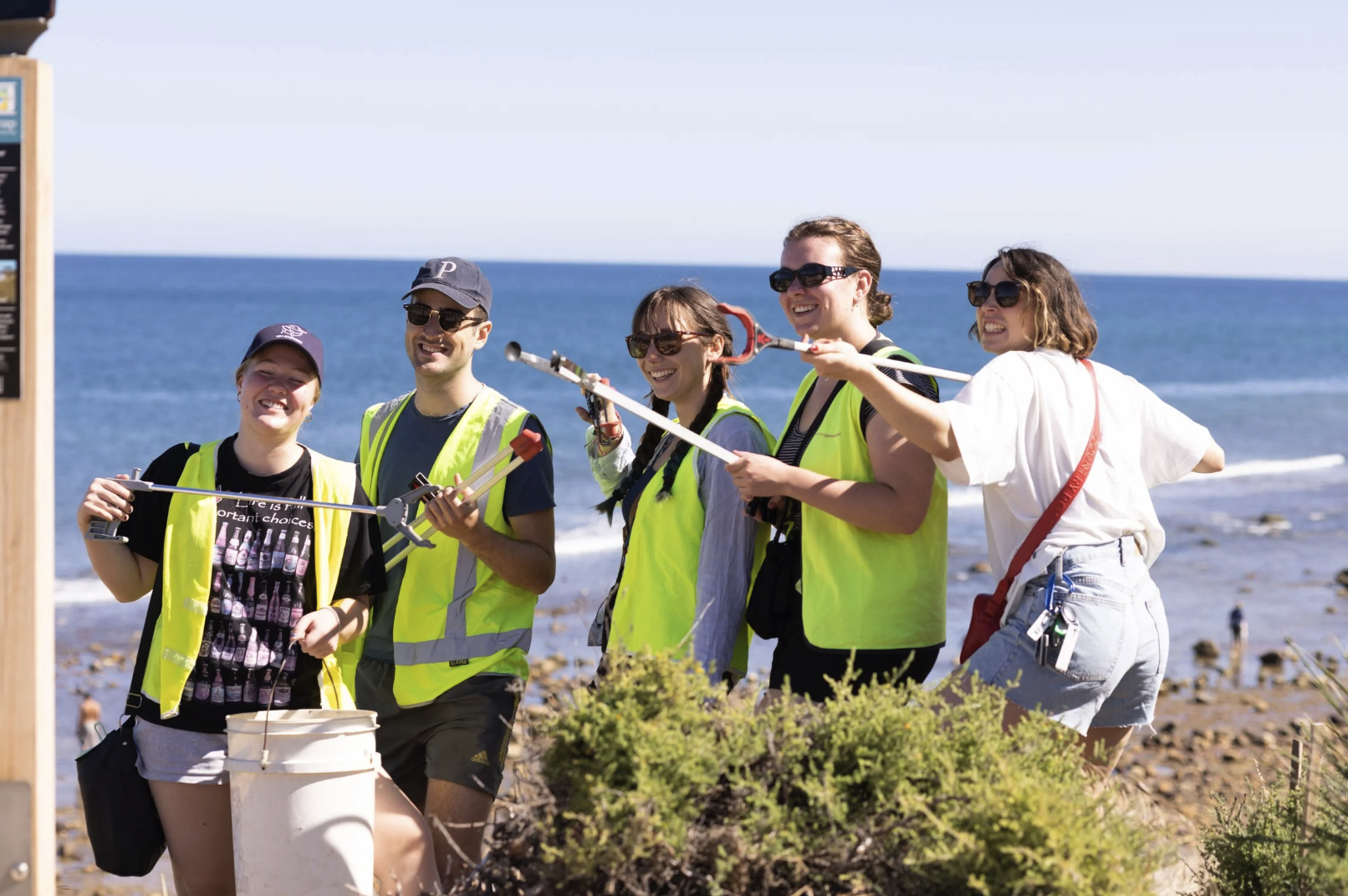
Marine scientists are collecting more underwater imagery than ever before. Autonomous vehicles, drop cameras, diver surveys and long-term monitoring programs are producing millions of images each year. The challenge is no longer collecting data. It is turning those images into usable management-ready information.
New research around the Squidle+ platform shows how that bottleneck is starting to shift.

Another shows how the same platform can be used to train machine learning models that classify key marine ecological indicators at national scale

A third paper detailing how Squidle+ is helping to build machine learning models to detect urchins, a key species threatening shallow reefs along eastern Australia
The bottleneck: too many images, not enough time
Squidle+ was built to deal with a practical problem. Marine imaging has expanded faster than scientists can efficiently analyse it. The platform brings together imagery management, annotation, map-based exploration, quality control and AI in one web-based system. It links to imagery stored in existing cloud repositories, which cuts down on duplication, and it allows different users and teams to work on the same datasets through shared workflows and permissions.
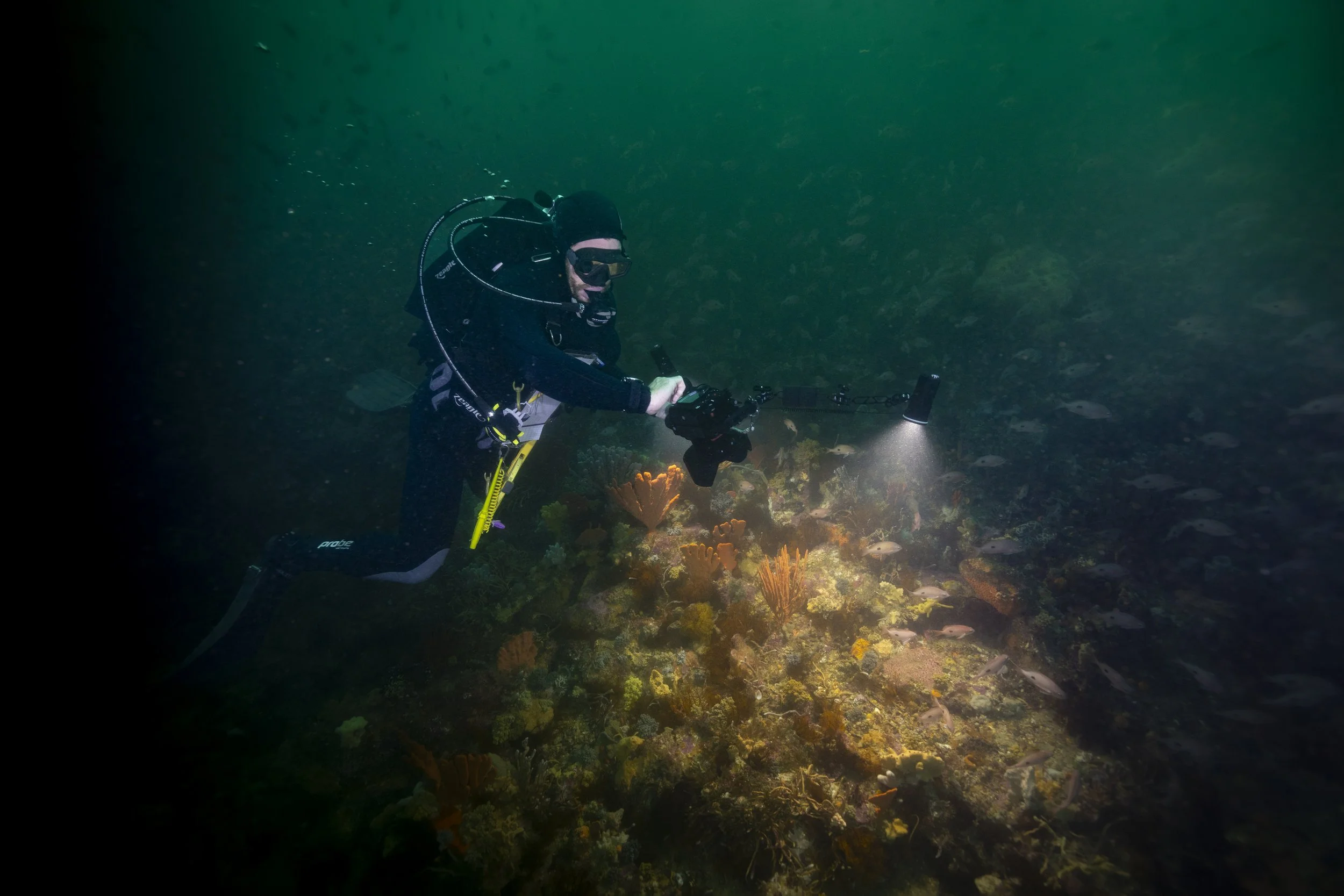

A key feature is the ability to translate between different annotation vocabularies, allowing datasets from different institutions or monitoring programs to be combined. That makes cross-project synthesis, broader reporting and large-scale machine learning training far more practical than it has been in the past. The platform paper also frames this as a major step toward making marine imagery more findable, accessible, interoperable and reusable.
Growing at Scale
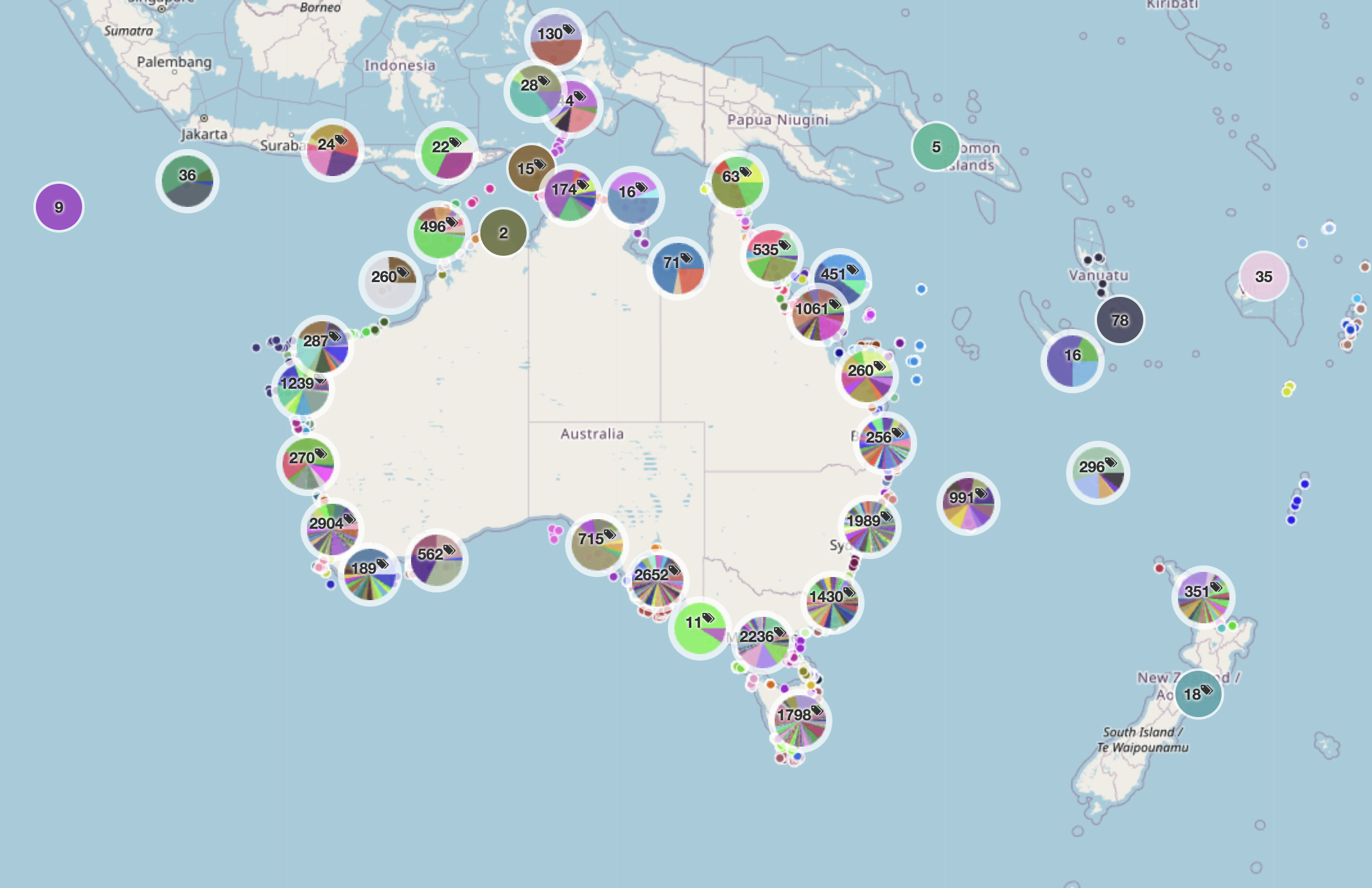
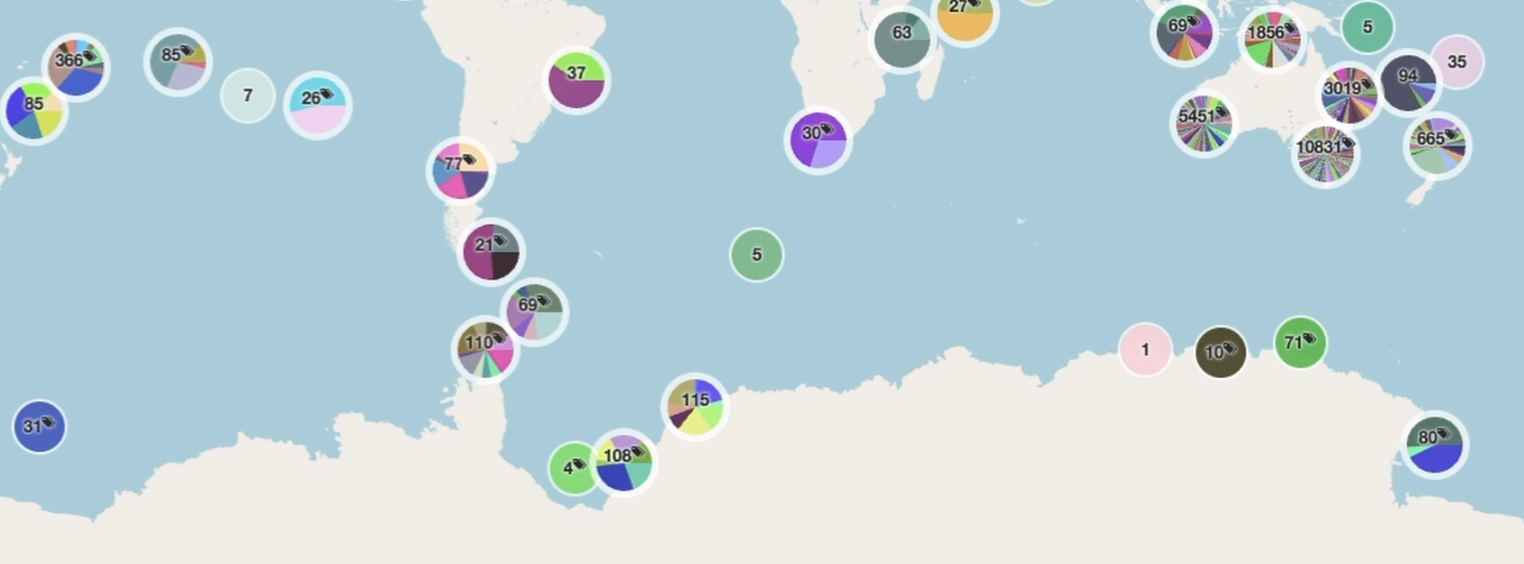
The scale of the platform is already substantial. As of April 2026, it had grown to
more than 10.5 million images
from 25,998 deployments across 1,147 campaigns
with more than 6.5 million annotations
produced by 2,586 registered users from 330 affiliations.
The platform paper describes it as the largest known repository of openly accessible georeferenced marine images with associated annotations.
A shared backbone for marine imagery
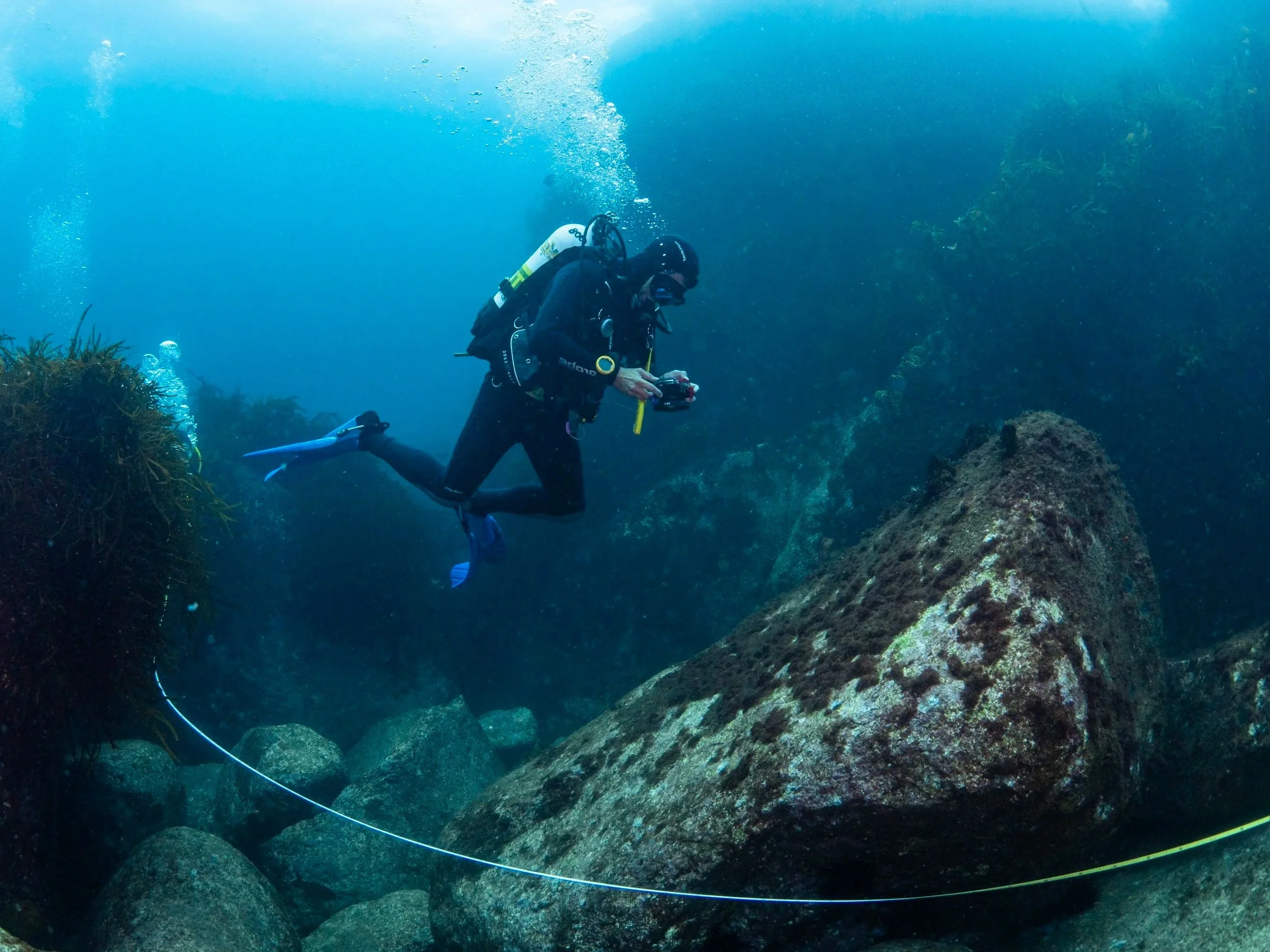

The platform is already being used across management, research, education and citizen science. Reef Life Survey, for example, used Squidle+ to coordinate photoquadrat annotation across a large volunteer network. According to the platform paper, that helped the program manually annotate more images in the first three years using Squidle+ than in the previous fifteen years combined with its earlier tools.
Squidle+ also provides the groundwork for machine learning. Large, well-labelled datasets are essential for training useful classifiers, and the platform’s architecture was built in a way that allows algorithms to sit alongside human annotators rather than outside the workflow.

In practice, Squidle+ acts as shared infrastructure for marine science. It supports collaborative annotation, quality control between people and algorithms, map-based data exploration, integration with other portals, and the export of training data for machine learning. This means imagery collected for one purpose can later support new research questions, monitoring programs or management decisions.
Training models at national scale
That shared infrastructure is what made the next step possible. Using Squidle+, researchers assembled a large and varied dataset of 1.7 million annotations across 150,000 images from 325 expeditions in Australian waters. That is a very different proposition from the smaller, single-project datasets often used to train image classifiers. By combining imagery from multiple campaigns, habitats and conditions, the researchers aimed to train models that could work across Australia’s coastline rather than only in one local setting.
The team built classifiers for three high-level habitat indicators, or Essential Ocean Variables: seagrass cover, canopy-forming macroalgae cover and hard coral cover.
These align with the kinds of variables prioritised by the Australian National Marine Science Committee and the Global Ocean Observing System for tracking ocean health and ecosystem change. The researchers also trained a species-specific classifier for Ecklonia radiata, a dominant kelp species across much of temperate Australia.
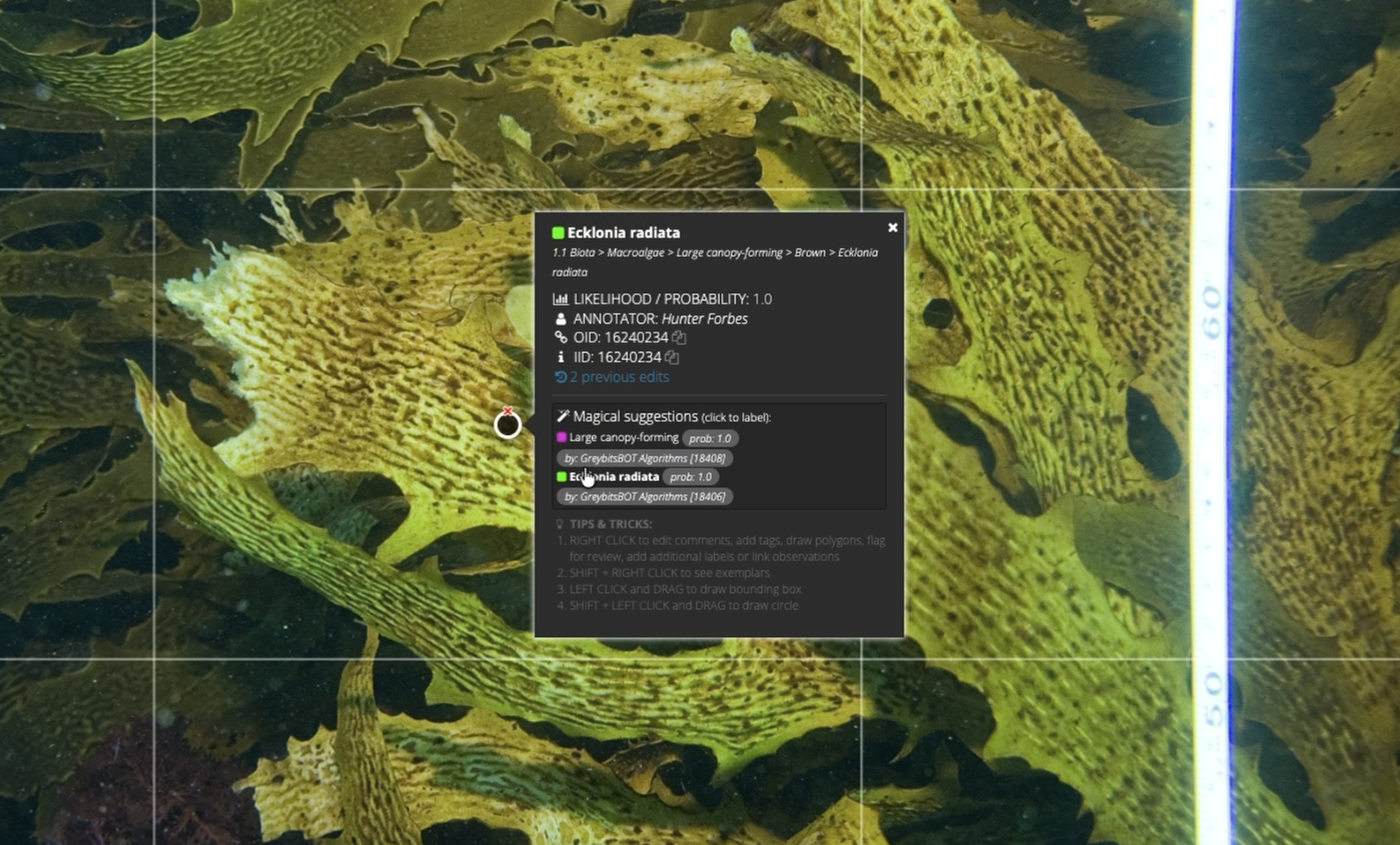
Turning Images Into Management-ready Insight
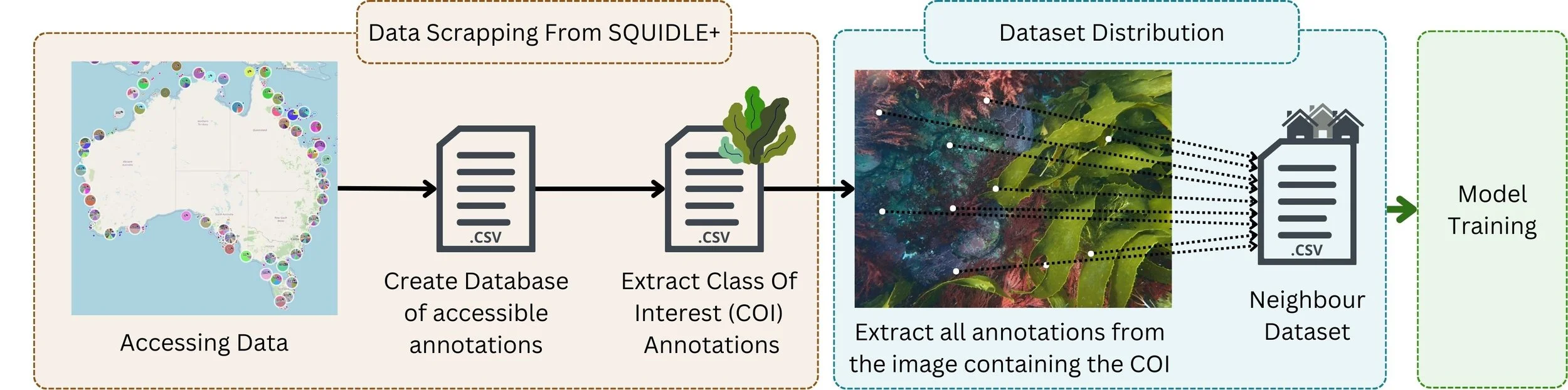
Because the training imagery in the habitat-classifier study came from many different sources, the researchers had to deal with variation in image resolution, survey design and visual conditions. To manage that, they introduced methods including adaptive cropping and a neighbour-distribution dataset strategy, designed to make this particular training dataset more useful across mixed image sets rather than assuming every campaign looked the same.

The availability of these classifiers in Squidle+ coupled with its efficient data export capabilities, allows platforms to seamlessly integrate large amounts of data into management-ready products. One example is the Reef Life Explorer, which uses data from Squidle+ for its reef health indicator “habitat-cover”. This represents canopy-forming macroalgae in temperate waters and can be viewed alongside other indicators of reef health globally.

The resulting classifiers achieved 85 to 93 per cent test accuracy for the habitat indicators and 96.5 per cent for Ecklonia radiata, with testing carried out on full campaign datasets that had been excluded from training. For the Great Southern Reef, that approach is especially relevant. Ecklonia radiata, a dominant canopy-forming seaweed across much of the reef, is a key habitat-former, and kelp cover can be used as an important indicator of ecosystem health and biodiversity. The strong performance of the Ecklonia and canopy-forming macroalgae models points to real potential for tracking kelp-dominated habitats more efficiently across large areas.
Squidle+ underpins Australia’s Integrated Marine Observing System’s (IMOS) Understanding Marine Imagery (UMI) initiative. IMOS is enabled by the National Collaborative Research Infrastructure Strategy (NCRIS).Detecting the destroyers: Urchinbot and the fight for kelp forests
While classifiers like the LeoBots models work at the level of habitat cover, what proportion of the seafloor is kelp, coral or seagrass, other ecological questions require knowing where specific animals are and how many of them are present. In kelp forest ecosystems, one of the most urgent of those questions concerns sea urchins. A new paper published in Marine Environmental Research introduces Urchinbot, an open-source machine learning model designed to detect and classify the key sea urchin species responsible for catastrophic kelp loss across northeastern New Zealand and southeastern Australia.
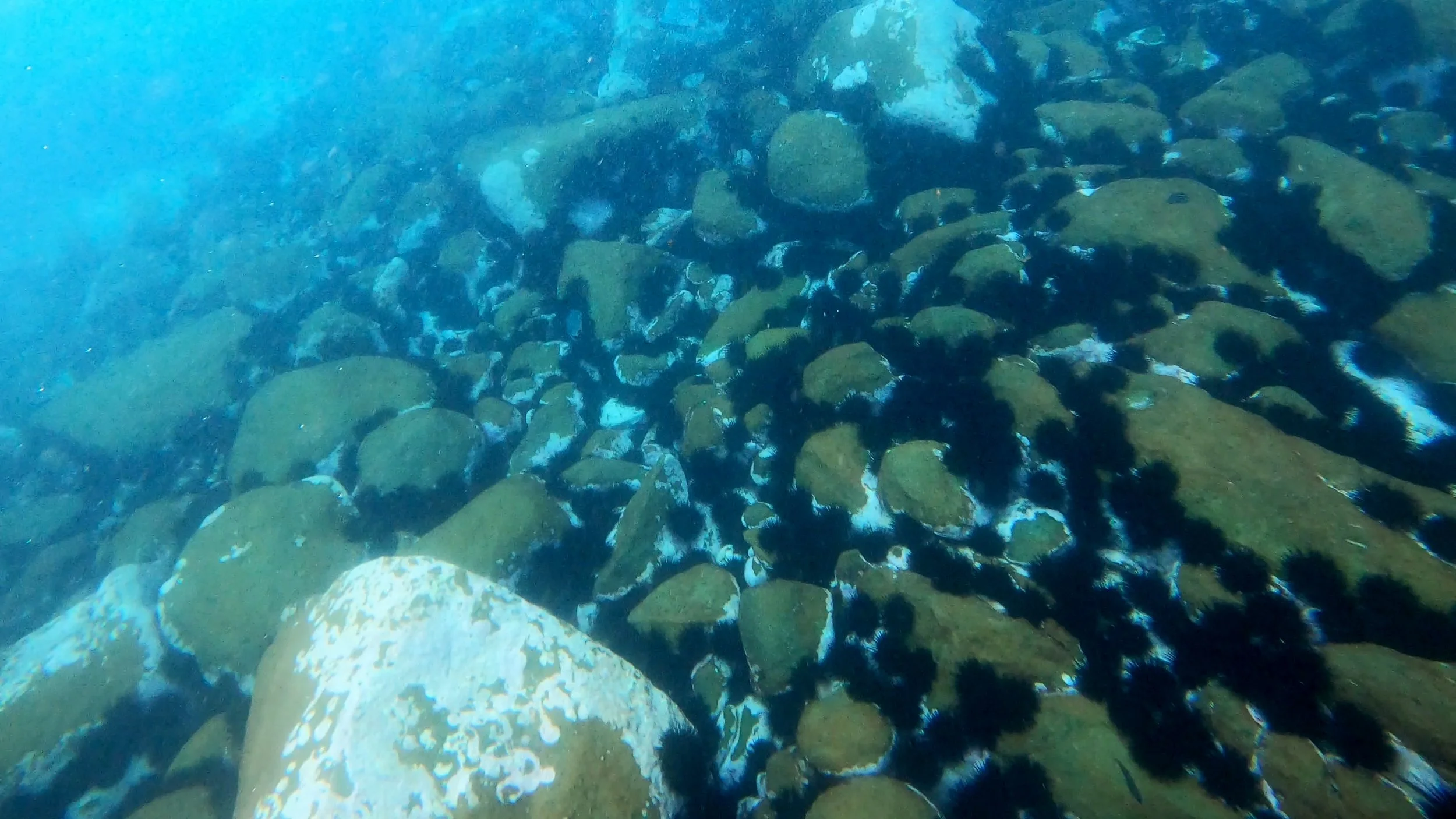

The ecological backdrop is serious. In northeastern New Zealand, an estimated 30 per cent of shallow reefs are now considered urchin barrens. In New South Wales, more than half of the nearshore rocky reefs along the central and southern coastline have already been reduced to that state. In Tasmania, modelling suggests that 50 per cent of kelp forests could be lost by 2030 without intervention.
Three sea urchin species drive much of this damage: Evechinus chloroticus in New Zealand, and Centrostephanus rodgersii and Heliocidaris erythrogramma across both countries. Understanding where these species are, in what numbers, and how their distributions are shifting is fundamental to any management response, but manual annotation of the underwater imagery needed to answer those questions has become a serious bottleneck. Thanks to the UrchinBot, researchers are now able to accurately process the 6,000-image northeastern New Zealand dataset and produce 33,000 individual annotations in less than two and a half hours.
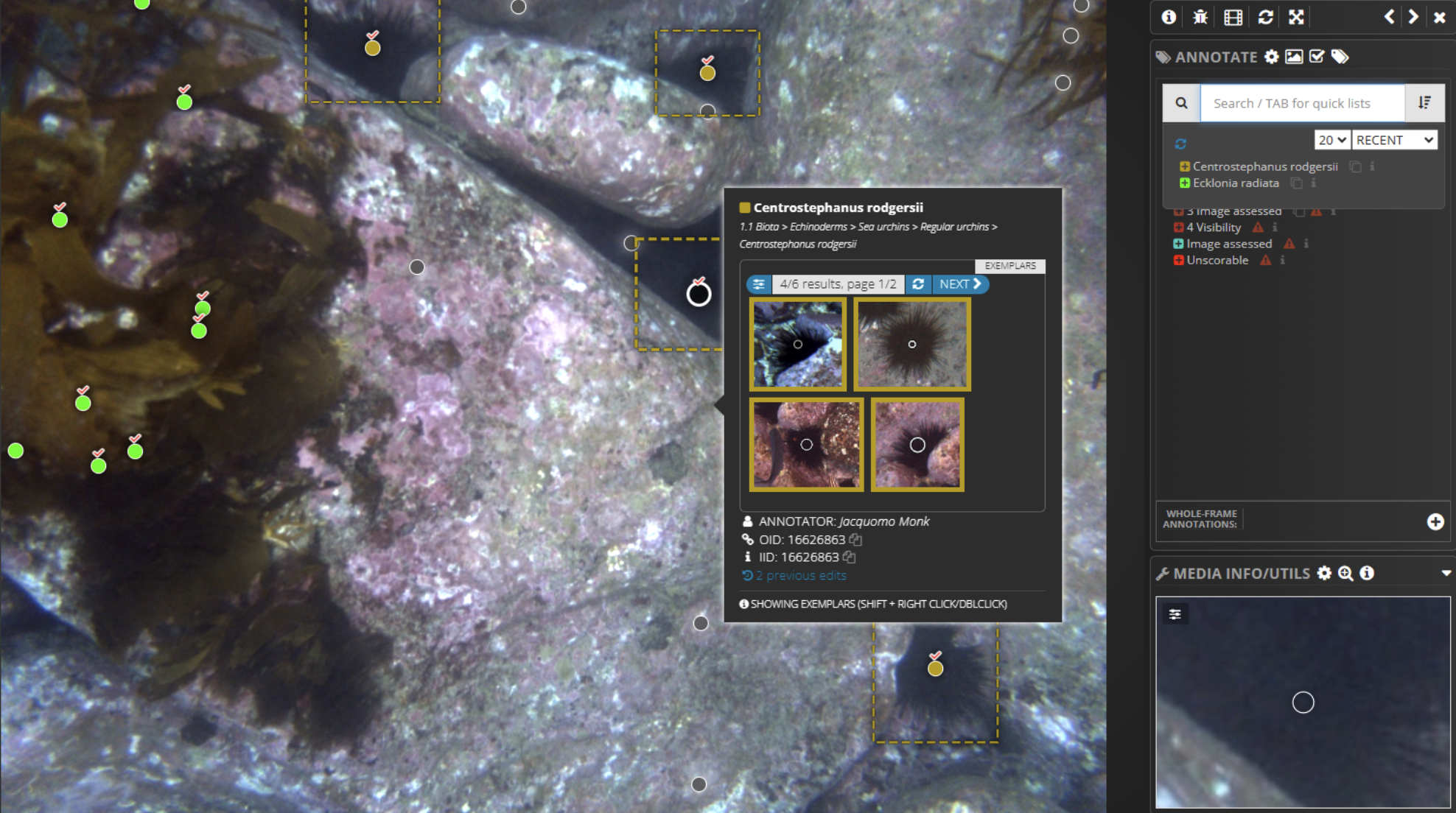
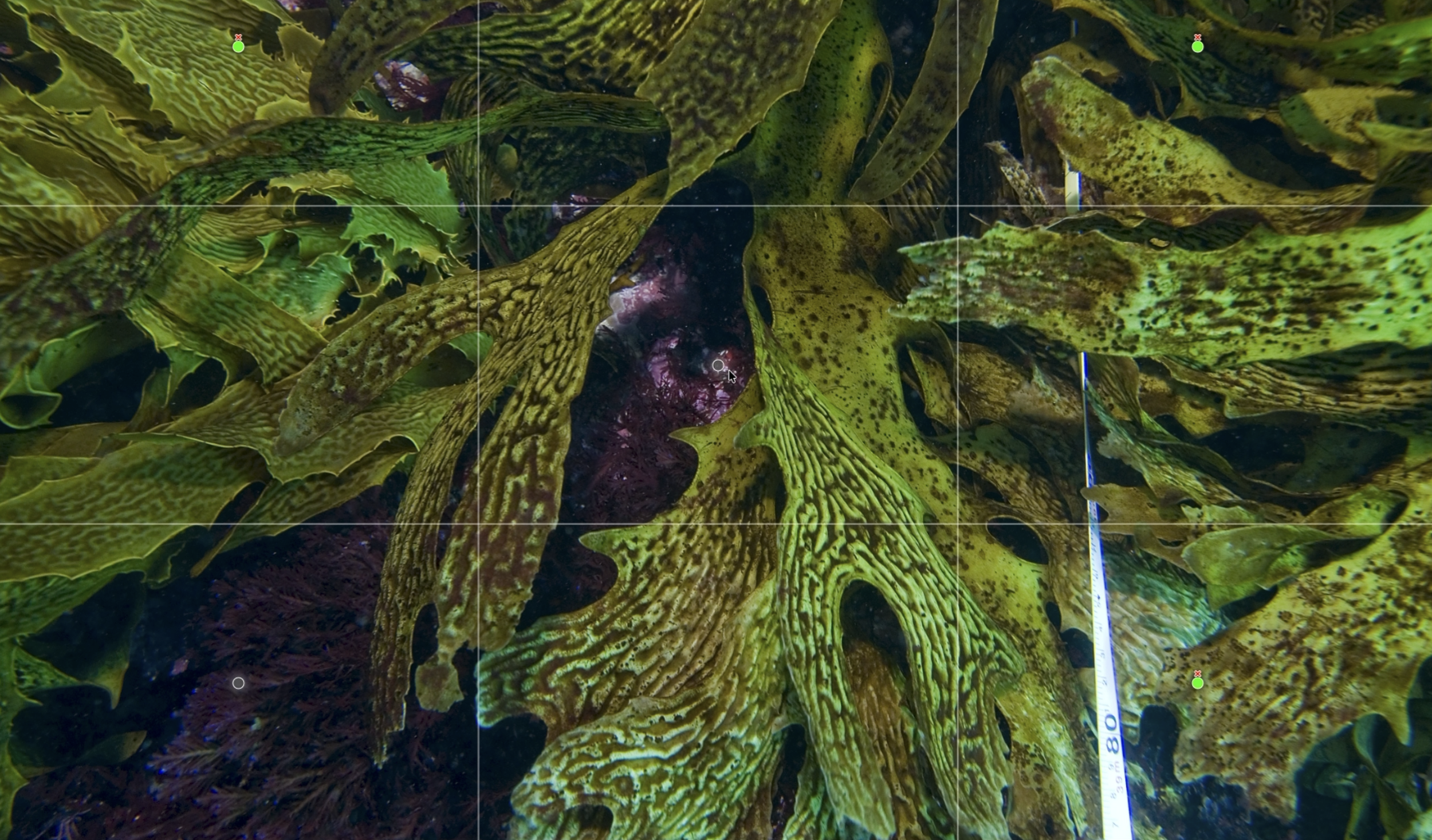
The Squidle+ annotation interface is displaying a partially annotated underwater image captured by the IMOS AUV Sirius at Elephant Rock, St Helens, Tasmania. Coloured points represent automated classifications, with tick symbols indicating human‑verified labels. A pop‑up window provides exemplar images of Centrostephanus rodgersii to support rapid confirmation of labels.Like the LeoBots classifiers, Urchinbot is openly available and fully integrated into Squidle+. Researchers can select imagery, run the model for automated annotation and then review or refine the results within the same platform, without needing their own computing infrastructure or specialist software. The source code is available on GitHub and the annotation datasets used for training and validation are archived through both Squidle+ and Zenodo. That combination of open code, open data, and platform integration is increasingly the model for how ecological AI tools reach the researchers who need them.
you may also like
CONNECT